Microscoop Mint Protein-Pickable Microscope
AI microscopy-guided photo-biotinylation
No antibody panels
Unbiased subcellular proteomics analysis
High resolution, high sensitivity and high specificity
Broad sample compatibility
Detail
Brief Introduction
Microscoop® Mint is a groundbreaking spatial proteomics platform that has been used to reveal novel protein constituents at specific subcellular regions of interest for many biological problems. Microscoop® Mint performs microscopic scooping, i.e. automated ultra-content microscopy-guided photo-biotinylation to photolabel and isolate/pick enough subcellular proteins for mass spectrometry-based proteomic discovery. It is an unprecedented spatial pulldown technology that enables unbiased subcellular proteomic identification in high resolution, high sensitivity, and high specificity.

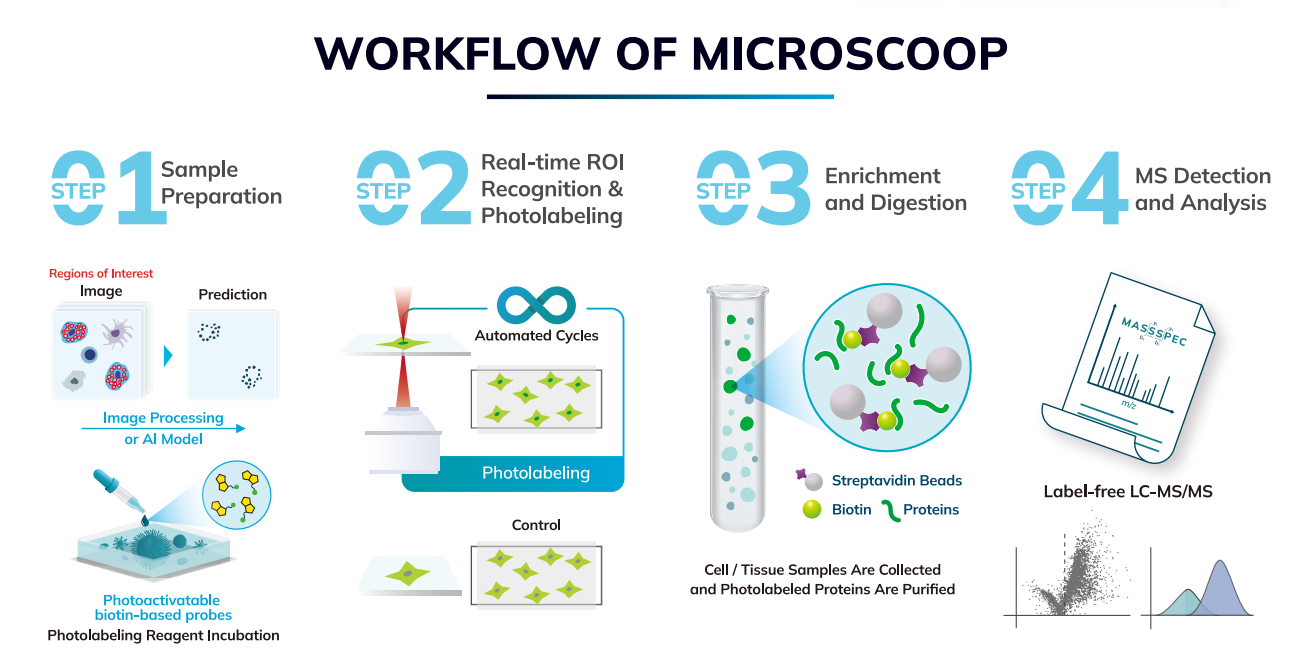
Workflow
STEP 1
REAL-TIME IMAGE ANALYSIS
Photolabeling kit (i.e. Synlight-RichTM Kit) is first added to a cell or tissue sample for a photochemical reaction.After the sample is loaded onto the stage, Microscoop® takes an image (or images of multiple colors) of the sample at one field of view (FOV) at a time. The image or images are analyzed in real time by Microscoop's software AutoscoopTM, which executes traditional image processing or Al deep learning to segment the user's region of interest. Pre- or post-processing can be included to enhance segmentation accuracy.
STEP 2
PATTERNED PHOTO-BIOTINYLATION
A femtosecond light source is controlled to illuminate the segmented region of interest one paime. This patterned illumination triggers targeted protein photo-biotinlyation in high spatial precision through the reactions of light-sensitive probes of Synlight-RichTM Kit. This patterned photolabeling is repeated for thousands of FOVs automatically to assure that enough proteins are biotinlyated for later proteomics analysis using mass spectrometry.
STEP 3
PROTEIN EXTRACTION
Photolabeled cells or tissues are scraped from the slide or chamber. Materials from multiple slides or chambers can be pooled together to increase the total protein contents.The scraped sample is then treated with reagents of protein extraction kit (i.e. SynpullTM Kit) to lyse the sample, enrich the proteins by immunoprecipitation, and digest them into peptides for proteomics analysis.
STEP 4
PROTEOMIC IDENTIFICATION
The collected peptides are sent to a mass spectrometer to perform LC-MS/MS analysis. Proteomes of both the photo-labeled and unlabeled (CTL) samples are obtained. By comparing the control and photolabeled proteomes, a location-specific proteome at the region of interest is obtained in high sensitivity, high specificity, and high spatial precision. Validation can be done by colocalization of immunostaining or additional functional assays.
Product Characteristics
1、Unbiased Discovery: Reveal the proteome without targeted panels;
2、High Precision: 25 nm labeling precision for subcellular labeling;
3、Superior Sensitivity: Discover low copy number proteins from increased dynamic range due to subcellular protein isolation
4、Broad Sample Compatibility: Use with FFPE/fresh frozen/fixed frozen tissue samples or fixed cells;
5、Sample Agnostic: No antibody panels, No molecular modifications, Native cell or tissue environment.
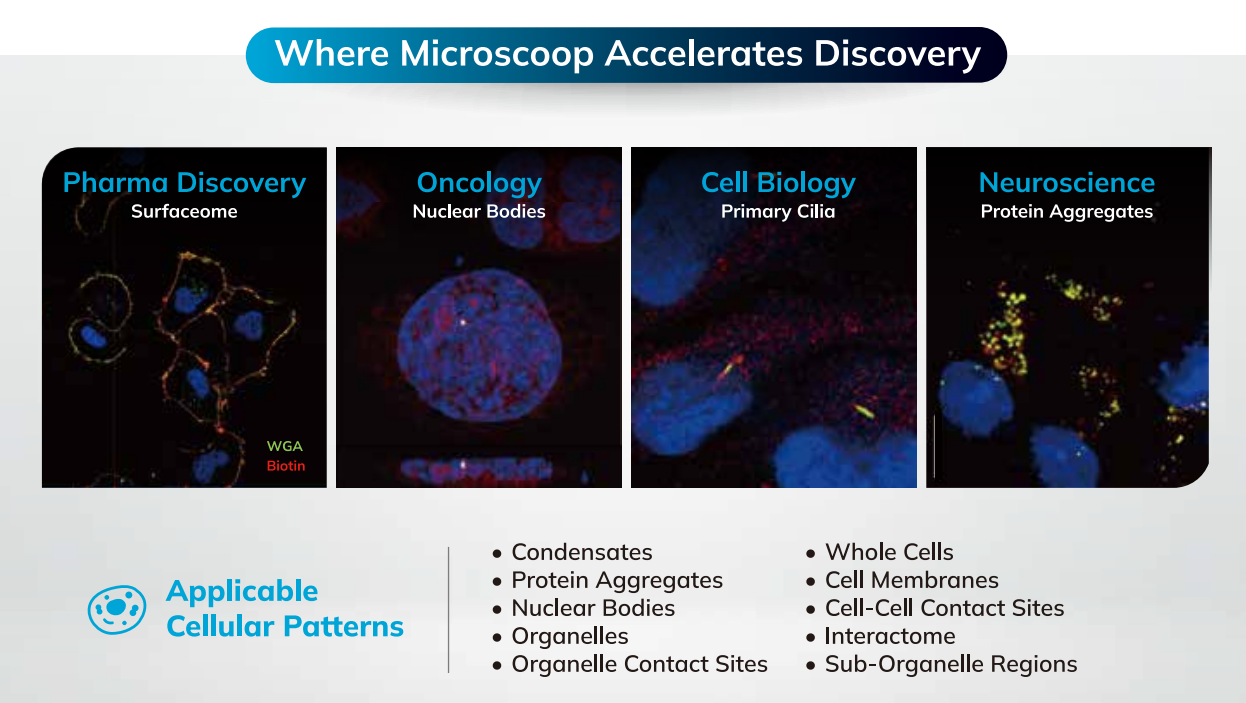
Broad Discovery Applications
Microscoop® Mint can be used for broad discovery applications, including: Neuroscience, Cancer Biology, Cell Biology, Immunology, Developmental Biology, Infectious Diseases, Aging, Metabolic Diseases, Inflammatory Diseases, Stem Cell Research, Immuno-Oncology, Pathology, Druggable Target Discovery, Biomarker Discovery and more.
Application
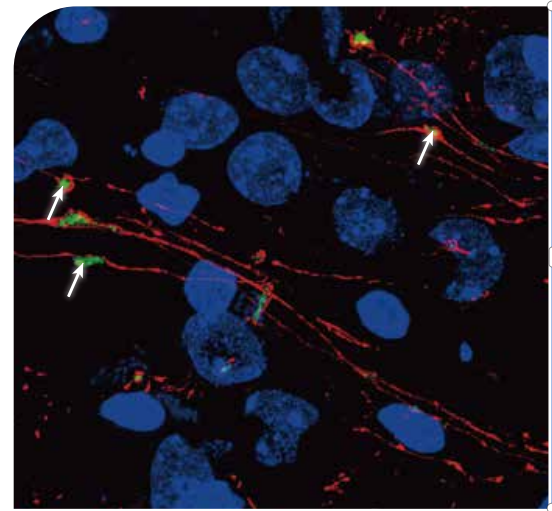
1、NEUROSCIENCE
Microscopy-recognizable structures such as β amyloids, reelin positive neurons at medical entorhinal layer Ⅱ, or dendritic spines can be studied using Microscoop-enabled spatial enrichment. Here is an example of hippocampal mossy fiber boutons photo-biotinylated by Microscoop® (Alexa488-neutravidin, green) for subsequent protein pulldown and proteome discovery.
2、CANCER BIOLOGY
Problem such as identifying E-cadherin-associated proteins of metastatic cells, proteins at the cancer cell-T cell inferface, and the proteome of Ki67+ cells in triple negative breast cancer can be addressed by Microscoop-enabled spatial enrichment and the subsequent proteomics analysis. Here is an example of the early-stage cancer marker cytokeratin19 around lesions in mouse pancreatic cancer.
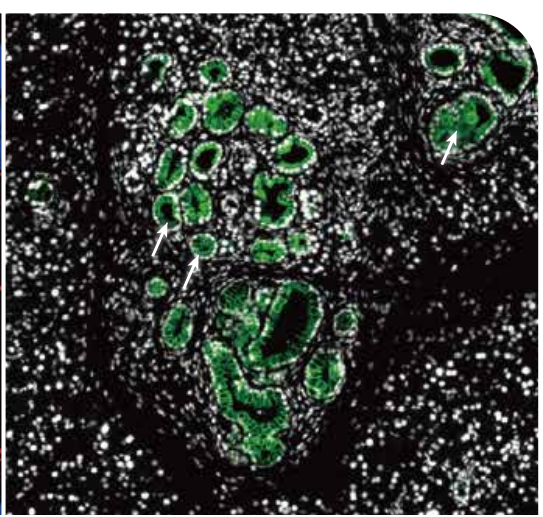
3、CELL BIOLOGY
Proteomes of subcellular features such as larger extracellular vesicles, stress granules, filopodia tips, focal adhesion, or ER-mitochondria interface can be identified using Microscoop® and mass spectrometry. Here shows an example of peroxisomes (PEX14, green) that can be photo-biotinylated using Microscoop® for subcellular spatial isolation and proteomics analysis.
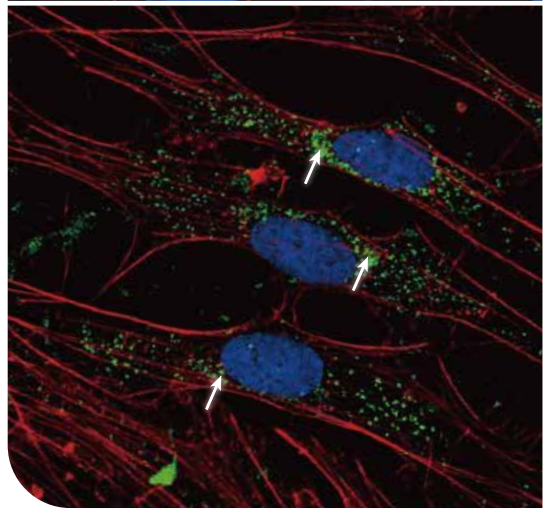
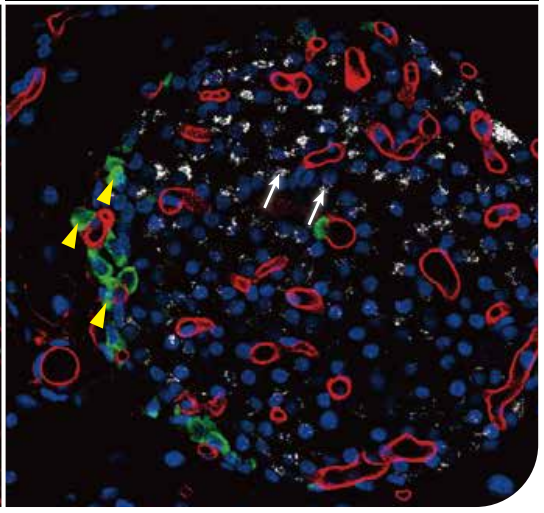
4、METABOLIC BIOLOGY
Microscoop-enabled proteomic discovery can be performed on other biological problems in immunology, metabolic diseases, developmental biology, infectious diseases, etc. Here is an image of pancreatic islet, where one can isolate α cells (glucagon, green), β-cells (insulin, white), or blood vessels (WGA, red) and identify novel protein constituents.